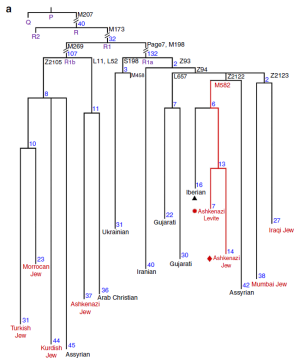
Over at Nature Communications, Rootsi et al. report on a newly discovered Ashkenazi-specific subclade of R1a, defined by the M582 mutation. They argue that it's a marker of Near Eastern origin, and based on the comprehensive data in their paper, I'd say they're correct. However, it's important to note that this doesn't preclude an ultimate Eastern European or Central Asian source of M582 in the Near East. For instance, an R1a mutation ancestral to M582 might have been introduced by the proto-Iranians from the steppe into what is now Iran during the early Indo-European dispersals. Indeed, that's actually what Figure 1a from the study suggests (phylogenetic tree of R1a below). The paper is open access, but here are a few quotes anyway:
Haplogroup R1a-M582 was only sporadically observed in Europe, the Diaspora residence of Ashkenazi Jews. Notably, it was not identified among 2,149 samples (including 922 R1a-M198) of non-Jews from East Europe, where the Ashkenazi Jewish community flourished in recent centuries (Table 1).
...
Within 1,068 West/North European samples (106 R1a-M198), M582 was observed in just one German sample, and among 3,756 Central/South European samples (710 R1a-M198), it was found only in one Hungarian and one Slovakian sample (Table 1).
...
Among 3,739 Near Eastern samples (303 R1a-M198), R1a-M582 was identified in various populations, with the highest frequency occurring within Iranians collected from the southeastern Kerman population who self-identified as Persians, northwestern Iranian Azeri and in Cilician Anatolian Kurds, at 2.86%, 2.50% and 2.83%, respectively (Table 1). In contrast, among 2,164 samples from the Caucasus (211 R1a-M198), R1a-M582 was found in just one Nogay sample (Table 1).
...
Considering the historical records of Ashkenazi Jews, three potential geographic sources should be considered: the Near East, which was the geographic location for the ancient Hebrews; Europe, which was the residence of the Ashkenazi Jewish Diaspora and the region in which they evolved for nearly two millennia; and the region overlapping with the no longer extant mid-11th Century Khazarian Khaganate, whose ruling class has been suggested to have converted to Judaism [18]. Our data render the latter source highly unlikely since the Khazarian Khaganate overlapped with the Northern Pontic-Caspian steppe and the North Caucasus region, in which just one Nogay sample carried the R1a-M582 haplogroup (Table 1). Furthermore, the Nogays, formerly a powerful Kipchak Turkic-speaking nomadic confederation, are relatively recent inhabitants of the Caucasus, and the STR haplotype of the sole R1a-M582 Nogay sample lies outside of the Levite cluster. Had the Caucasus region been the source for the Ashkenazi modal lineage, we likely would have found R1a-M582 Y-chromosomes in some of its 20 local populations examined in our sample of more than 2,000 Y-chromosomes (Table 1).
...
Near Eastern populations are the only populations in which haplogroup R1a-M582 was found at significant frequencies (Table 1). Moreover, the representative samples displayed substantial diversity even within this geographic region (Fig. 1b). Higher frequencies and diversities often suggest lineage autochthony.
Citation...
Rootsi, S. et al. Phylogenetic applications of whole Y-chromosome sequences and the Near Eastern origin of Ashkenazi Levites. Nat. Commun. 4:2928 doi: 10.1038/ncomms3928 (2013).
See also...
The Poltavka outlier
He was short (<160 cm), probably lactose intolerant, had an exceedingly long melon (cranial index = 72.6), and belonged to mtDNA haplogroup K2a. In other words, he was a typical Neolithic farmer, and clearly different from the average modern-day inhabitant of the North European Plain.
No doubt, his people were largely replaced by newcomers from the east and also west during the frequent population shifts in the region after the Neolithic (see here). However, the stable isotope analysis suggests that he ate a lot of millet, which is known as a typically Slavic cultigen in Europe.
ABSTRACT: In 2007 a ceremonial complex representing the Globular Amphora Culture was discovered in Kowal (the Kuyavia region, Poland). Radiocarbon dating demonstrated that the human remains associated with the complex are of similar antiquity, i.e. 4.105 ± 0.035 conv. and 3.990 ± 0.050 conv. Kyrs. After calibration, this suggests a period between 2850 and 2570 BC (68.2% likelihood), or more specifically, 2870 to 2500 BC (95.4% likelihood). Morphological data indicate that the skeleton belonged to a male who died at 27–35 years of age. The unusual morphology of his hard palate suggests this individual may have had a speech disorder. Stable oxygen isotope values of the individual's teeth are above the locally established oxygen isotope range of precipitation, but due to sample limitations we cannot conclusively say whether the individual is of non-local origin. Stable carbon and nitrogen isotope ratios were analyzed to reconstruct the diet of the studied individual, and show a terrestrial-based diet. Through ancient DNA (aDNA) analysis, the mtDNA haplogroup K2a* and lactose intolerance as evidenced by homozygous C-13910 allele were identified. These aDNA results are the first sequences reported for an individual representing the Globular Amphora Culture, enriching the still modest pool of human genetic data from the Neolithic.
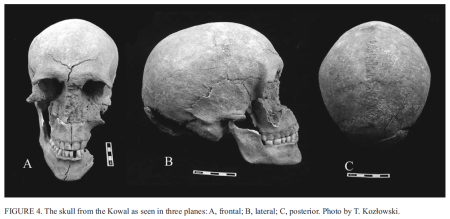
Kozłowski T., Stepańczak B., Laurie J. Reitsama, Osipowicz G., Szostek K., Płoszaj T., Jędryhowska-Dańska K., Pawlyta J., Paluszkiewicz C., Witas H.W. Osteological, chemical and genetic analyses of the human skeleton from a Neolithic site representing the Globular Amphora Culture (Kowal, Kuyavia region, Poland), Anthropologie [In Press]
See also...
Polish "Goths" enjoyed their millet, while Polish "Vikings" did not
One of the most enduring myths or cliches concerning European genetic structure is that Poles carry Mongolian admixture. This claim has been repeated so often that it's now regarded by many as fact, including at academic level. Poles apparently acquired this admixture during Mongol and Turkic raids on Eastern Europe during the Middle Ages.
For a long time it was impossible to verify or debunk such claims due to a lack of genetic data from European and Asian populations at high enough resolution. However, that's no longer a problem. Indeed, I've run a wide range of detailed analyses as part of my Eurogenes Genetic Ancestry Project which have shown that my Polish sample does not carry inflated levels of Asian ancestry relative to other Northern and Central Europeans. But in this post it's probably more useful for me to focus on results from peer-reviewed studies to make my point.
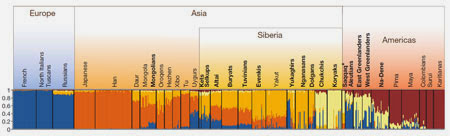
I'll start with a look at genome-wide genetic data (aka. autosomal DNA). The bar plot below was featured in a recent study on genetic substructures in European Russia (1). It shows the results of an ADMIXTURE analysis at K=2 (two ancestral populations assumed) using 52,808 SNP markers, which splits the genomes of the samples into West and East Eurasian components. As you'd expect, these components are modal in Europeans and Han Chinese, respectively. This northern Han Chinese sample from Beijing is certainly a very useful proxy for Mongolians, because the East Eurasian component it creates is basically the same one that peaks in the least admixed Mongolian samples in other studies (for example, see 2).
The lowest levels of the East Eurasian component are carried by Latvians, Poles, Germans, Czechs and Italians. But this looks like noise anyway, because the Chinese carry a reciprocal amount of the West Eurasian component. In other words, it's most likely not a real signal of recent admixture from East Asia, but shared prehistoric Eurasian ancestry. If it's not noise, and actually represents admixture from an East Asian source like the Mongolians, then it's difficult to explain why it appears at a clearly higher level in Italians than Poles.
A couple of the Czechs do carry inflated amounts of the East Eurasian component, and I'd say they're Czech Roma with significant South Asian admixture who weren't removed as outliers from the dataset. It's also worth mentioning that the Komi, Finns and Russians (like the Rus_HGDP sample from Kargopol) show higher levels of this component than the Central Europeans. However, this doesn't necessarily mean they have Mongolian ancestry. Indeed, another study has shown that the Kargopol Russians carry various Siberian-specific components, rather than the type of East Asian influence which makes up the majority of Mongolian genome-wide genetic structure (2).
Moreover, the Mongols never raided North Russia or Finland, where in fact these components peak in Europe today. So the most plausible explanation for the relatively high levels of Siberian admixture there is Finno-Ugric or Uralic ancestry. Indeed, the Finns obviously speak a Finno-Ugric language, and so do some North Russian groups, while many others did until recently.
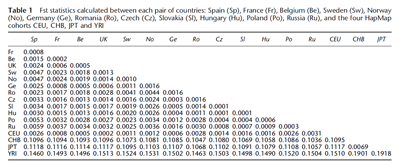
Below is a different kind of analysis of autosomal DNA. It's a pairwise Fixation Index (Fst) test between 16 global populations based on 129,673 SNPs (3). It includes two East Asian samples, the same Han Chinese set as in the above ADMIXTURE analyses, and a Japanese sample from Tokyo. The results appear to correlate very closely with geography, which suggests that we're basically looking at the effects of isolation-by-distance. Interestingly, the Polish sample from Lodz and Warsaw (Po) is genetically more distant from the Han Chinese (CHB) and Japanese (JPT) than are the Swedes (Sw), Norwegians (No) and Germans (Ge). The differences aren't big, but they're consistent, which means that at the very least these Poles can't be carrying more East Asian ancestry than the Scandinavian and German samples.
Next up is a global Principal Component Analysis (PCA) from the same study, featuring the same samples and markers. Again, it's another way of looking at variation in autosomal DNA, and again the results indicate that Poles don't show any special genome-wide genetic links to East Asians. The Polish samples are sitting close to the top of the European cluster and overlap strongly with other Northern and Central Europeans. Indeed, many of the outliers pulling towards the West African (YRI) and East Asian (CHB + JPT) clusters are French, British and German. These individuals are possibly carrying very recent non-European ancestry due to colonial links between Western Europe and the third world. There are also a considerable number of Russians streaming towards the East Asians, and that's probably due to the Finno-Ugric mediated Siberian influence in North Russia described above.
 It might also be useful to assess the level of West Asian or Near Eastern autosomal admixture in Poland compared to other parts of Europe, because many of the Turkic groups which ended up on the Eastern European steppe were largely of West Asian origin. Therefore, if there was a significant genetic contribution from such groups to the Polish gene pool, then Poles today ought to show inflated affinity to West Asian populations compared to other Central and Northern Europeans. The simple answer is that they don't, which can be clearly seen on the pairwise Fst clustering analysis below based on approximately 101,000 SNPs (4).
Y-chromosome or paternal markers tell the same story as autosomal DNA. The most common Y-DNA haplogroup in Poland is R1a1a, making up about 50% of all the Y-chromosome lineages in the country. Based on academic and commercial testing of hundreds of Polish samples to date, it's safe to say that the most dominant subclades of this Eurasian haplogroup in Poland are the European-specific R-M458 and R-Z280 (5, 6). These subclades are closely related to the Scandinavian-specific R-Z284, because all three markers come off the Z283 branch of the R1a1a haplotree. The Z283 mutation is also mostly confined to Europe and possibly originated there.
The main Asian-specific subclade of R1a1a, known as R-Z94, appears to be extremely rare in Poland. This is the subclade that makes up the main share of the R1a1a in Mongolian and Turkic populations of Central Asia, as well as in South Asians. However, even the very few R-Z94 cases among Poles are more closely related to Ashkenazi lineages than those of Mongolians or Central Asians. What all of this means, of course, is that Mongolian or Turkic paternal ancestry isn't "hiding" in Poland under all that R1a1a.
The most common Y-DNA haplogroup among Mongolians and their close ethnic kin is C3. There's actually a particular lineage of C3 called the "star-cluster chromosome" which pops up regularly in purported descendents of the infamous Mongolian warlord Genghis Khan. This might mean that it's a marker of paternal ancestry from Genghis himself, which has been suggested in several academic papers (7, 8, 9). In any case, the haplotype is considered to be very young, probably less than 1000 years, and widespread in areas of Asia where the Mongol hordes were most active during the early Middle Ages. Therefore, it seems to be an excellent signal of Mongolian genetic influence from this key period. Below is a map of its frequencies in a variety of Asian groups.
It might also be useful to assess the level of West Asian or Near Eastern autosomal admixture in Poland compared to other parts of Europe, because many of the Turkic groups which ended up on the Eastern European steppe were largely of West Asian origin. Therefore, if there was a significant genetic contribution from such groups to the Polish gene pool, then Poles today ought to show inflated affinity to West Asian populations compared to other Central and Northern Europeans. The simple answer is that they don't, which can be clearly seen on the pairwise Fst clustering analysis below based on approximately 101,000 SNPs (4).
Y-chromosome or paternal markers tell the same story as autosomal DNA. The most common Y-DNA haplogroup in Poland is R1a1a, making up about 50% of all the Y-chromosome lineages in the country. Based on academic and commercial testing of hundreds of Polish samples to date, it's safe to say that the most dominant subclades of this Eurasian haplogroup in Poland are the European-specific R-M458 and R-Z280 (5, 6). These subclades are closely related to the Scandinavian-specific R-Z284, because all three markers come off the Z283 branch of the R1a1a haplotree. The Z283 mutation is also mostly confined to Europe and possibly originated there.
The main Asian-specific subclade of R1a1a, known as R-Z94, appears to be extremely rare in Poland. This is the subclade that makes up the main share of the R1a1a in Mongolian and Turkic populations of Central Asia, as well as in South Asians. However, even the very few R-Z94 cases among Poles are more closely related to Ashkenazi lineages than those of Mongolians or Central Asians. What all of this means, of course, is that Mongolian or Turkic paternal ancestry isn't "hiding" in Poland under all that R1a1a.
The most common Y-DNA haplogroup among Mongolians and their close ethnic kin is C3. There's actually a particular lineage of C3 called the "star-cluster chromosome" which pops up regularly in purported descendents of the infamous Mongolian warlord Genghis Khan. This might mean that it's a marker of paternal ancestry from Genghis himself, which has been suggested in several academic papers (7, 8, 9). In any case, the haplotype is considered to be very young, probably less than 1000 years, and widespread in areas of Asia where the Mongol hordes were most active during the early Middle Ages. Therefore, it seems to be an excellent signal of Mongolian genetic influence from this key period. Below is a map of its frequencies in a variety of Asian groups.
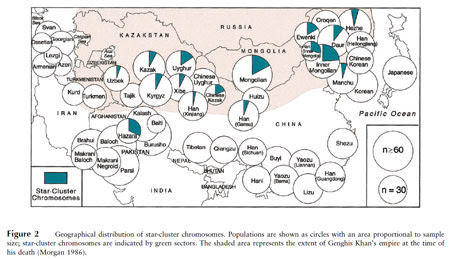 However, this marker has never been recorded in Poland, where C3 itself is extremely unusual. Indeed, Derenko et al. conclude that based on the paucity of the "star-cluster haplotype" among ethnic Russians, the Mongol hordes of the Middle Ages didn't even leave a genetic imprint on European Russia.
However, this marker has never been recorded in Poland, where C3 itself is extremely unusual. Indeed, Derenko et al. conclude that based on the paucity of the "star-cluster haplotype" among ethnic Russians, the Mongol hordes of the Middle Ages didn't even leave a genetic imprint on European Russia.
It is known that the Mongol Empire expanded over a considerable part of Eastern Europe by 1248 due to the khan Batu’s conquests. Russian principalities were vassal states of the Mongol Empire until 1480. However, we found no genetic traces of the Mongol sovereignty over Russia (in the form of male lineages of the cluster of Genghis Khan descendants) in the Russian population.
Poles do sporadically carry East Eurasian-specific mtDNA (ie. maternal) lineages, and it was recently suggested by Mielnik-Sikorska et al. that some of these lineages (ie. C4a1a, G2a and D5a2a1a1) might "possibly reflect relatively recent contacts of Slavs with nomadic Altaic peoples" (10). However, the authors also note that these markers have a wide distribution across Eurasia and might actually represent prehistoric gene-flow between Asia and Europe.
In fact, ancient DNA results have recently revealed unexpected frequencies of East Eurasian-specific mtDNA haplogroups - including C1, C4a2, C5, D* and Z1a - in Northern and Eastern European remains from the Neolithic and Mesolithic (11, 12, 13, 14). The samples were always small, but nevertheless it's useful to note that the incidence of the eastern mtDNA lineages was much higher in these ancient samples than in any modern Eastern European populations west of the Volga. What this suggests is that the vast majority of East Eurasian ancestry in Europe might have arrived there thousands of years before the Mongol incursions, and much of it has been lost since then, rather than gained, due to continuous population movements from west to east across the continent.
To add another twist to the tale, it actually seems that all Europeans do show significant prehistoric East Eurasian genome-wide admixture, so perhaps it's only the ancient eastern mtDNA markers that have almost gone extinct? The topic has been now been discussed in a couple of papers, and apparently this admixture shows higher affinity to modern Amerindians than North or East Asians (15), which is very curious indeed. But the details are sketchy because the results so far have been based on DNA from modern samples, so hopefully we can learn more very soon thanks to analyses of genome-wide markers from prehistoric Eurasian remains. In any case, a group of Poles was featured in one of these papers, and this is how they compared to other Europeans in a formal mixture test with the ADMIXTOOLS software (16).
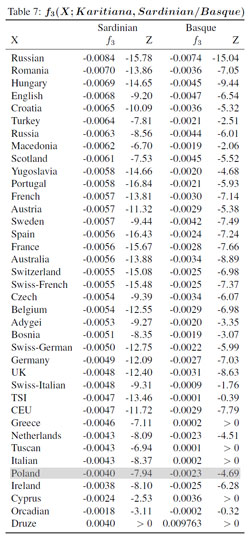 The positions of the samples in the table are based on the "Sardinian" f3-statistic, which indicates the strength of the Amerindian-like admixture signal when Karitiana Indians and Sardinians are chosen as references. The results appear fairly random in some cases, and that's probably because they're skewed by such factors as genetic drift and sample size - for instance, heavy drift might dampen the signal, while a larger sample size might increase it. But the outcomes are also clearly influenced by relatively recent admixture from Siberia, Central Asia and/or the Indian subcontinent. That's because the samples known to carry such admixtures, like the HGDP Russians and Turks, are at the top of the table. Therefore, it's worth noting that the Polish sample is found at the bottom, which is in line with results from all the other analyses presented above.
Please note that the information in this post pertains to the current Polish population by and large, and to individuals with all grandparents of ethnic Polish origin. It's not relevant to individuals whose recent ancestors came from within the borders (or former borders) of Poland but their ethnic origins were uncertain. Keep in mind that Poland was not as ethnically or genetically homogenous before World War II as it is today. Also worth noting is that in a country of almost 40 million people, some individuals will have very atypical pedigrees, and it might even be possible to find someone with, say, Papuan ancestry in Poland if we look hard enough.
By the way, I've no idea who made that meme at the top of the post, or where it was published originally, but I think it's very appropriate here. Please let me know if it violates any copyright laws and I'll take it down.
References...
1. Khrunin AV, Khokhrin DV, Filippova IN, Esko T, Nelis M, et al. (2013) A Genome-Wide Analysis of Populations from European Russia Reveals a New Pole of Genetic Diversity in Northern Europe. PLoS ONE 8(3): e58552. doi:10.1371/journal.pone.0058552
2. Morten Rasmussen et al., Ancient human genome sequence of an extinct Palaeo-Eskimo, Nature 463, 757-762 (11 February 2010) doi:10.1038/nature08835; Received 30 November 2009; Accepted 18 January 2010
3. Simon C Heath et al, Investigation of the fine structure of European populations with applications to disease association studies, European Journal of Human Genetics (2008) 16, 1413–1429; doi:10.1038/ejhg.2008.210
The positions of the samples in the table are based on the "Sardinian" f3-statistic, which indicates the strength of the Amerindian-like admixture signal when Karitiana Indians and Sardinians are chosen as references. The results appear fairly random in some cases, and that's probably because they're skewed by such factors as genetic drift and sample size - for instance, heavy drift might dampen the signal, while a larger sample size might increase it. But the outcomes are also clearly influenced by relatively recent admixture from Siberia, Central Asia and/or the Indian subcontinent. That's because the samples known to carry such admixtures, like the HGDP Russians and Turks, are at the top of the table. Therefore, it's worth noting that the Polish sample is found at the bottom, which is in line with results from all the other analyses presented above.
Please note that the information in this post pertains to the current Polish population by and large, and to individuals with all grandparents of ethnic Polish origin. It's not relevant to individuals whose recent ancestors came from within the borders (or former borders) of Poland but their ethnic origins were uncertain. Keep in mind that Poland was not as ethnically or genetically homogenous before World War II as it is today. Also worth noting is that in a country of almost 40 million people, some individuals will have very atypical pedigrees, and it might even be possible to find someone with, say, Papuan ancestry in Poland if we look hard enough.
By the way, I've no idea who made that meme at the top of the post, or where it was published originally, but I think it's very appropriate here. Please let me know if it violates any copyright laws and I'll take it down.
References...
1. Khrunin AV, Khokhrin DV, Filippova IN, Esko T, Nelis M, et al. (2013) A Genome-Wide Analysis of Populations from European Russia Reveals a New Pole of Genetic Diversity in Northern Europe. PLoS ONE 8(3): e58552. doi:10.1371/journal.pone.0058552
2. Morten Rasmussen et al., Ancient human genome sequence of an extinct Palaeo-Eskimo, Nature 463, 757-762 (11 February 2010) doi:10.1038/nature08835; Received 30 November 2009; Accepted 18 January 2010
3. Simon C Heath et al, Investigation of the fine structure of European populations with applications to disease association studies, European Journal of Human Genetics (2008) 16, 1413–1429; doi:10.1038/ejhg.2008.210
4. Esko et al., Genetic characterization of northeastern Italian population isolates in the context of broader European genetic diversity, European Journal of Human Genetics advance online publication 19 December 2012; doi: 10.1038/ejhg.2012.229
5. Peter A Underhill et al., Separating the post-Glacial coancestry of European and Asian Y chromosomes within haplogroup R1a, European Journal of Human Genetics advance online publication 4 November 2009; doi: 10.1038/ejhg.2009.194
6. Family Tree DNA R1a1a and Subclades Y-DNA Project
7. Derenko et al., Distribution of the Male Lineages of Genghis Khan’s Descendants in Northern Eurasian Populations, Russian Journal of Genetics, 2007, Vol. 43, No. 3; DOI: 10.1134/S1022795407030179
8. Zerjal et al., The Genetic Legacy of the Mongols, Am J Hum Genet. 2003 March; 72(3): 717–721. PMCID: PMC1180246
9. Abilev S. et al., The Y-chromosome C3* star-cluster attributed to Genghis Khan's descendants is present at high frequency in the Kerey clan from Kazakhstan, Hum Biol. 2012 Feb;84(1):79-89. doi: 10.3378/027.084.0106.
10. Mielnik-Sikorska M, Daca P, Malyarchuk B, Derenko M, Skonieczna K, et al. (2013) The History of Slavs Inferred from Complete Mitochondrial Genome Sequences. PLoS ONE 8(1): e54360. doi:10.1371/journal.pone.0054360
11. Zsuzsanna Guba et al., HVS-I polymorphism screening of ancient human mitochondrial DNA provides evidence for N9a discontinuity and East Asian haplogroups in the Neolithic Hungary, Journal of Human Genetics advance online publication 15 September 2011; doi: 10.1038/jhg.2011.103
12. Alexey G Nikitin et al., Mitochondrial haplogroup C in ancient mitochondrial DNA from Ukraine extends the presence of East Eurasian genetic lineages in Neolithic Central and Eastern Europe, Journal of Human Genetics advance online publication, 7 June 2012; doi:10.1038/jhg.2012.69
13. Lillie, Malcolm C et al., Prehistoric populations of Ukraine: Migration at the later Mesolithic to Neolithic transition, Population Dynamics in Prehistory and Early History (2012), Publication Date: July 2012, ISBN: 978-3-11-026630-6, DOI: 10.1515/9783110266306.93
14. Der Sarkissian C, Balanovsky O, Brandt G, Khartanovich V, Buzhilova A, et al. (2013) Ancient DNA Reveals Prehistoric Gene-Flow from Siberia in the Complex Human Population History of North East Europe. PLoS Genet 9(2): e1003296. doi:10.1371/journal.pgen.1003296
15. Lipson et al., Efficient moment-based inference of admixture parameters and sources of gene flow, arXiv:1212.2555v1 [q-bio.PE]
16. Patterson et al., Ancient Admixture in Human History, Genetics: Published Articles Ahead of Print, published on September 7, 2012 as 10.1534/genetics.112.145037
See also...
R1a and R1b from an early Mongolian tomb